
Residue conservation of key positions in SWEET protein family
Residue conservation in prokaryotes

1 Residue numbers correspond to the structure of semiSWEET from Leptospira biflexa (PDB ID: 4QNC). For each residue, we have also provided the generic structure-based numbering scheme which conveys its relative position within a transmembrane helix with respect to the most conserved residue in that segment.
2 For details of residues that form extracellular and intracellular gates, selectivity filter and substrate-binding site, see references.
3 Only those residues which show conservation at least 15% are mentioned in the table.
4 Small and weakly polar residues: Ala, Gly, Ser, Thr and Cys; Hydrophobic: Leu, Ile, Val and Met.
Residue conservation in plants and metazoans
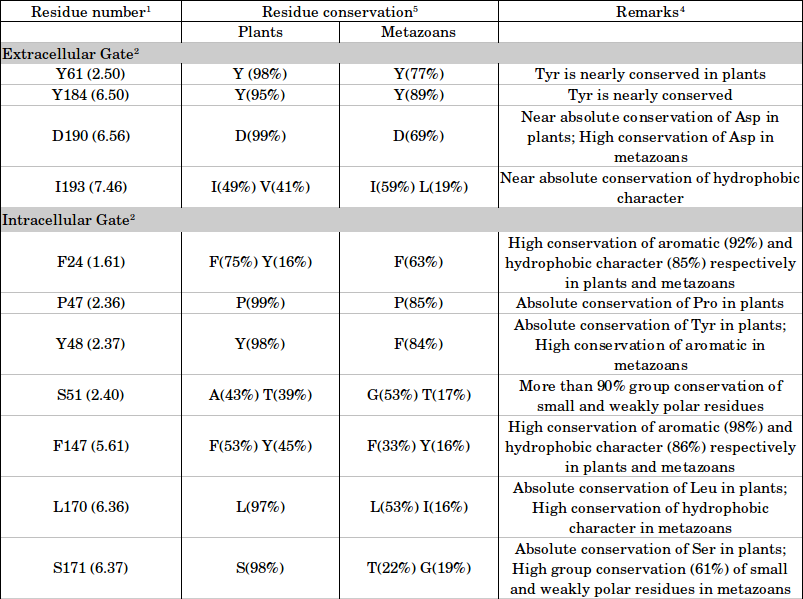
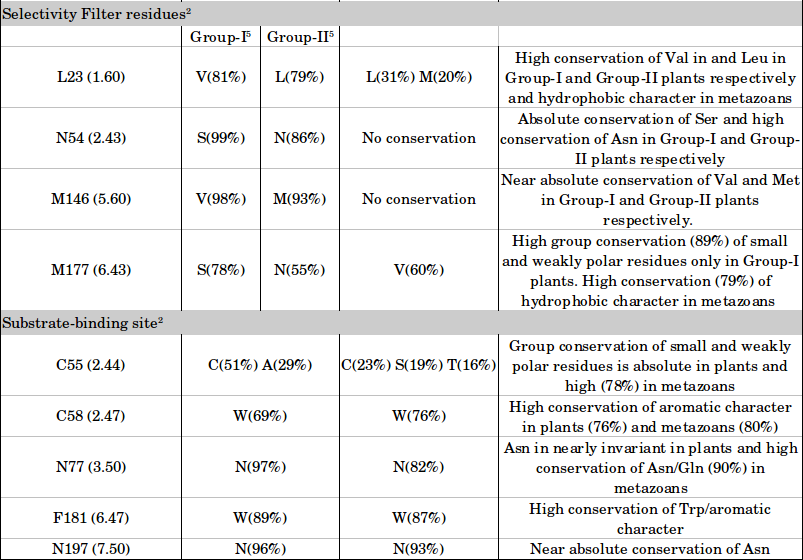
1 Residue numbers correspond to the structure of SWEET from Oryza sativa (PDB ID: 5CTG). For each residue, we have also provided the generic structure-based numbering scheme which conveys its relative position within a transmembrane helix with respect to the most conserved residue in that segment.
2 For details of residues that form extracellular and intracellular gates, selectivity filter and substrate-binding site see references.
3 Only those residues which show conservation at least 15% are mentioned in the table.
4 Small and weakly polar residues: Ala, Gly, Ser, Thr and Cys; Hydrophobic: Leu, Ile, Val and Met.
Group conservation of small and weakly polar residues
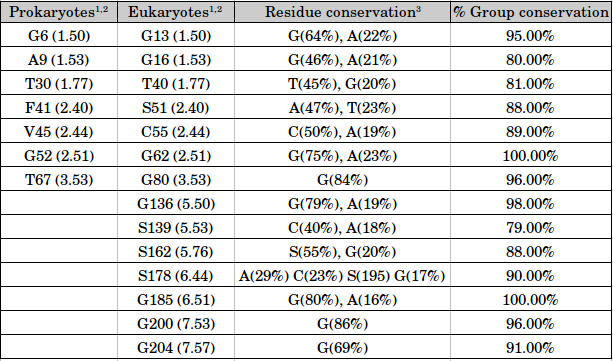
1 Residue numbers correspond to the semiSWEET structure from Leptospira biflexa (PDB ID: 4QNC) for prokaryotes and structure of SWEET from Oryza sativa (PDB ID: 5CTG) for eukaryotes.
2 For each residue, the generic structure-based numbering scheme is provided in brackets.
3 Only those residues which show conservation at least 15% are mentioned in the table.
References
- Molecular mechanism of substrate recognition and transport by the AtSWEET13 sugar transporter.Lei Han, Yongping Zhu, Min Liu, Ye Zhou, Guangyuan Lu, Lan Lan, Xianping Wang, Yongfang Zhao, and Xuejun C. Zhang. Pubmed:28878024
- Functional role of oligomerization for bacterial and plant SWEET sugar transporter family. Yuan Hu Xuan, YiBingHu, Li-Qing Chen, Davide Sosso, Daniel C. Ducat, Bi-Huei Hou, and Wolf B. Frommer. Pubmed:24027245
- Structure of a eukaryotic SWEET transporter in a homo-trimeric complex. Yuyong Tao, Lily S. Cheung, Shuo Li, Joon-Seob Eom, Li-Qing Chen, Yan Xu, Kay Perry, Wolf B. Frommer and Liang Feng. Pubmed:26479032
- Structural basis for the facilitative diffusion mechanism by SemiSWEET transporter.Yongchan Lee, Tomohiro Nishizawa, Keitaro Yamashita, Ryuichiro Ishitani, and Osamu Nurekia. Pubmed:25598322