
Average pairwise sequence identity of SWEET homologs from prokaryotes and eukaryotes and Generic numbering scheme for SWEET family of proteins
Pairwise global sequence alignment was performed between every SWEET and SemiSWEET homologs. The average sequence identity between and within the taxanomical groups(archaea, bacteria, plants and metazoans) SWEET homologs was derived.
The Triple Helix Bundles(THBs) - 1 and 2 of eukaryotic SWEET homologs were separated to compare with prokaryotic SemiSWEET homologs.
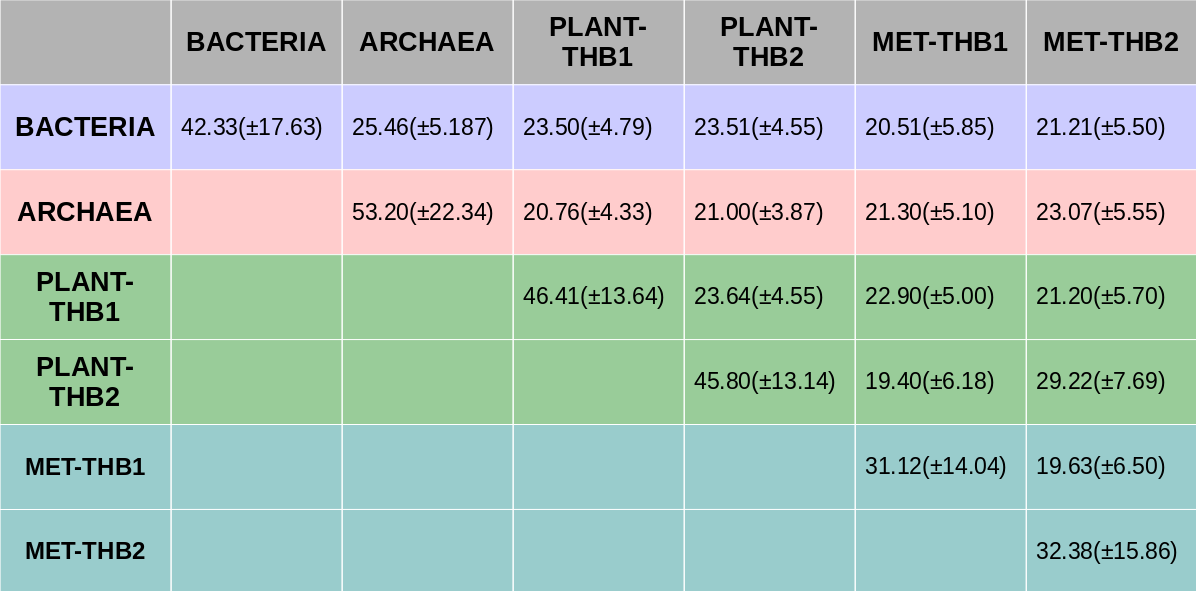
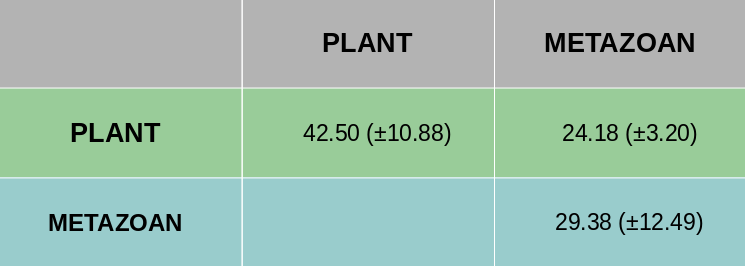
Average pairwise sequence identity between full length metazoan and plant SWEET proteins

The SWEET family of sugar transporters is very diverse, hence we have proposed a generic numbering scheme for SWEET family of proteins. We identified highly conserved residues in each transmembrane segment of semiSWEET and SWEET family members. The residue which is the highest conserved residue in a transmembrane segment is assigned as 50. Positions of other residues are relative to this residue. Residue 2.50 implies the most conserved residue in the second transmembrane segment. The position 2.47 refers the residue which is separated from 2.50 by three residues towards N-terminus. Positions of highly conserved residues in each transmembrane segment of semiSWEETs and SWEETs are given below. A similar numbering scheme is proposed for the superfamilies of GPCRs [1] and Major Intrinsic Proteins (MIPs) [2].
Generic Numbering scheme for SWEET family:
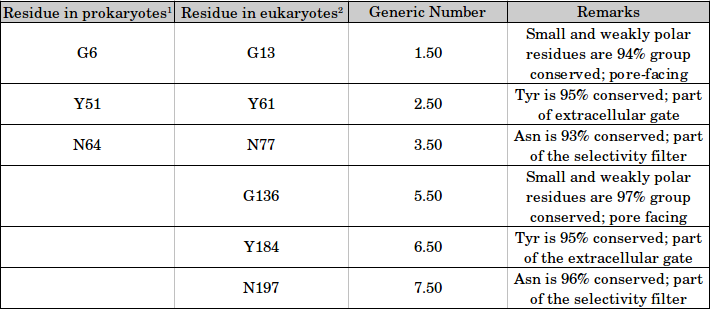
[1] Residue numbers according to the PDB ID 4QNC [2] Residue numbers according to PDB ID 5CTG
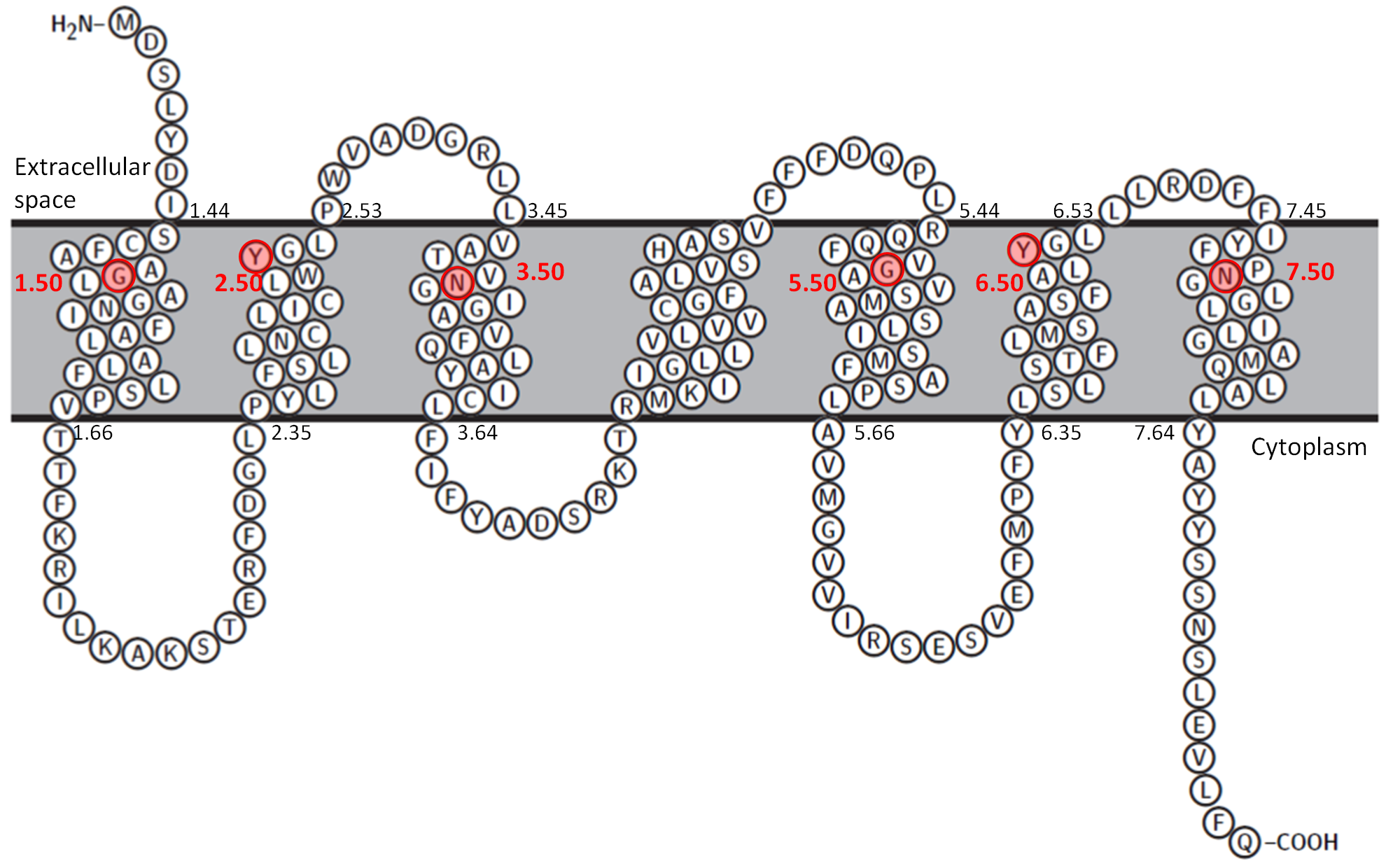
Snake diagrams showing the transmembrane regions SWEET proteins. The highly conserved residues in each transmembrane segment are shown in the red background. They are assigned a value of 50 and other residues within a transmembrane segment are numbered relative to this residue.
Two dimensional representation for rice SWEET homologue (PDB 5CTG)
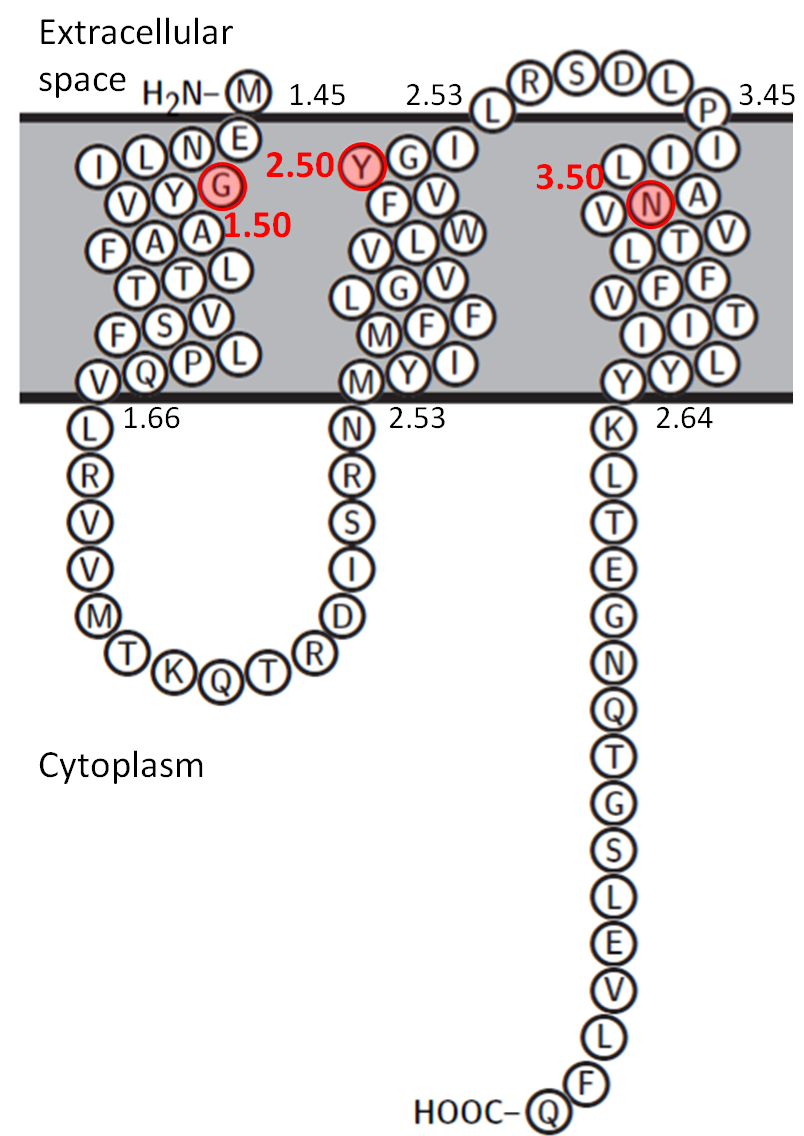
Two dimensional representation for bacterial SemiSWEET homologue (PDB 4QNC)
References
- Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Juan A.Ballesteros and Harel Weinstein. Methods in Neuroscience. 1995.
- Major intrinsic protein superfamily: channels with unique structural features and diverse selectivity filters. Ravi Kumar Verma, Anjali Bansal Gupta, and Ramasubbu Sankararamakrishnan. Methods Enzymol. 2015. Pubmed:25950979